Research
pH-dependent conformational dynamics of influenza hemagglutinin and the associated membrane-fusion process
Influenza virus hemagglutinin (HA) is a homotrimeric viral envelope glycoprotein that undergoes pH-triggered conformational changes in order to facilitate the fusion of the viral envelope with the
endosomal membrane of the infected host cell. The trimeric structure at neutral pH is made up a central helical stem formed by the three HA2 chains, with the three HA1 chains surrounding the HA2
chains. At low pH, HA1 moves away from HA2, which then undergoes conformational changes to form a needle-shaped structure exposing the N-terminal fusion peptide, while remaining connected to the
viral envelope through the C-terminus. The HA2 then bends at a hinge region to form a hairpin shaped structure, thus bringing the viral envelope very close to the endosomal membrane. While this model
of membrane fusion is well established, atomistic details of the pH-mediated conformational changes are poorly understood.
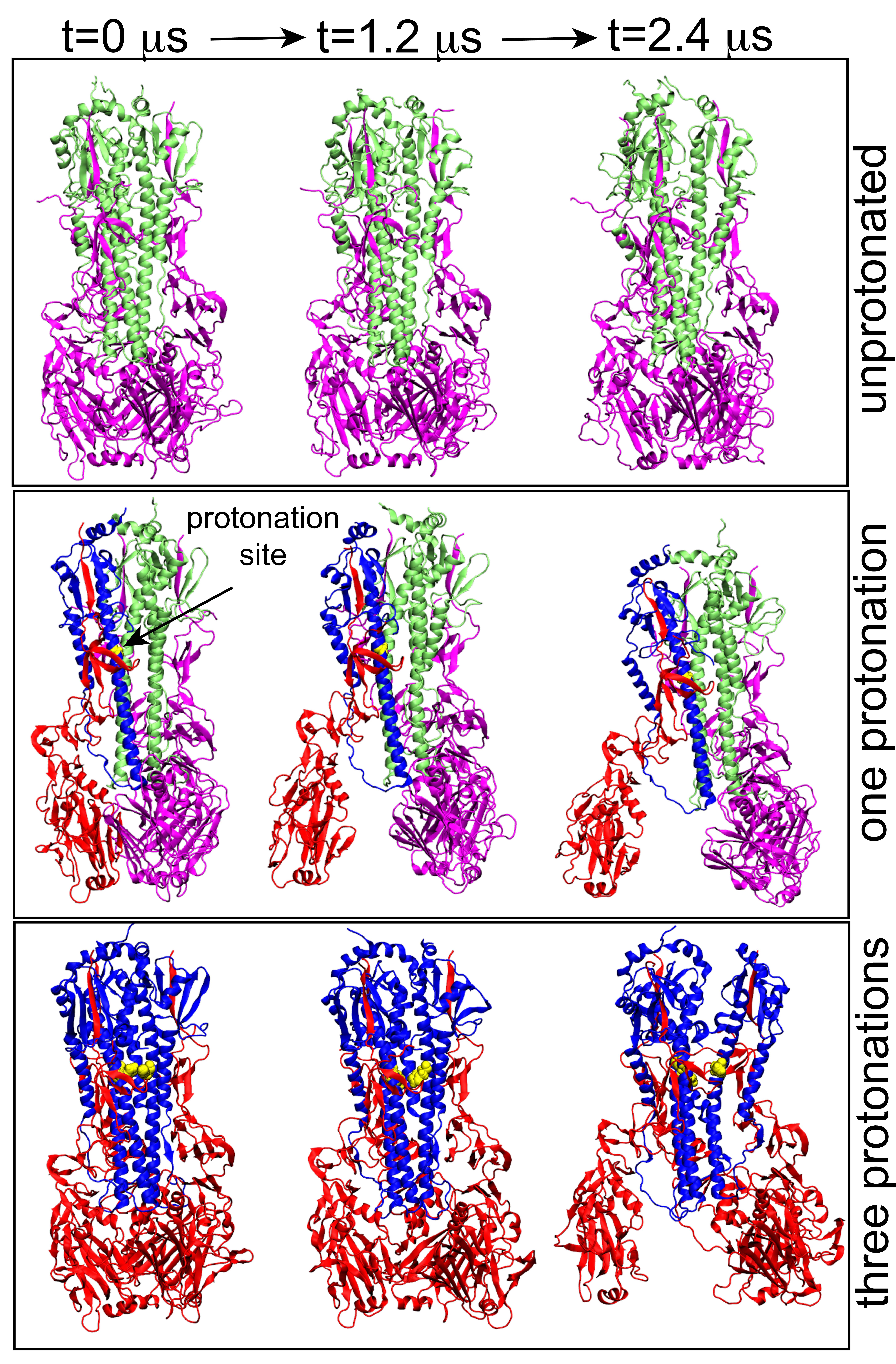
The HA2 hinge region includes a conserved histidine (H106). We believe that protonation of this conserved histidine at low pH (4.8-6.0) could be the trigger for the conformational changes associated
with membrane fusion. All-atom microsecond-level equilibrium MD simulations have been used to study the effects of protonating a single conserved histidine on HA conformational dynamics. The preliminary
data based on three 2.4 microsecond simulations reveals that the HA protein undergoes very large conformational changes even when a single histidine residue is protonated. No mechanistic framework at a
molecular level has been proposed for the coupling of conformational changes in influenza HA with histidine protonation. The initial results show that the protonation of H106 in the HA2 hinge region is
sufficient to cause the HA1 domains to move away from the HA2 domains over a time period of 2 microseconds. This was observed when H106 was protonated on all 3 HA2 monomers and on a single HA2 monomer.
On the other hand, when H106 was not protonated, the HA1 domains continued to sequester the HA2 domains.
|


